Protein structures, similarly to sequences, can be compared either globally, to reveal distant evolutionary relationships, or locally, to highlight similarities of specific group of residues such as enzyme active sites or ligand binding sites. Indeed two proteins can have similar functions even if their overall structures are completely different since the function depends on the position and identity of a small number of residues.
Superpose3D allows you to uncover these small but crucial similarities between protein structures. You can use it to study ligand binding sites, enzyme active sites, protein/protein interaction surfaces etc. The main advantage of Superpose3D is that it allows you to specify in a completely flexible way how the structure is represented during the computation (see the Examples).
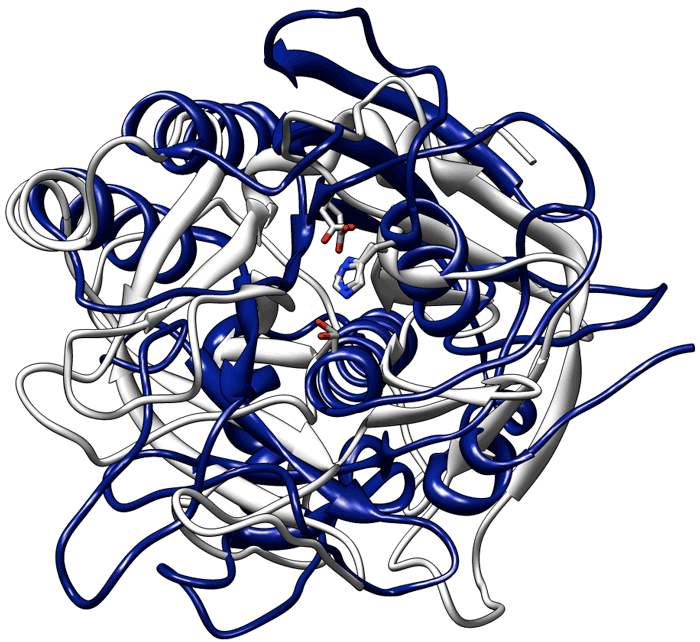
Local structural comparison programs are the ideal tool to discover instances of convergent evolution in protein structures (i.e. independent “inventions” of the same functional sites). The figure below depicts the most famous of such cases. The proteins trypsin (white) and subtilisin (blue) have completely different structures. However their active sites harbor an identical triad of catalytic residues (represented in stick in the figure). Indeed these two enzymes perform exactly the same reaction using the same catalytic mechanism!